Until 2007 there were no published reports of highly successful methods for culturing Cannabis sativa. Hemp was successfully cultured in China but at a success rate of a mere 85%. Then Ole Miss published “Thidiazuron-induced high-frequency direct shoot organogenesis of Cannabis sativa L.” in The Society for In Vitro Biology which detailed how they achieved a 95% success rate.
We found this paper and started making the medium. We were able to get great results with the vegetative propagation medium which utilizes hormones TDZ and gibbrellic acid. We used this medium to multiply and expand our library. The rooting medium calls for half strength MS salts and the exchange of hormones to only IBA and with the addition of activated charcoal. We had problems initially with the medium being watery after sterilization but we have since linked the issue to our gelling agent and have moved forward.
Synthetic seeds
During our research we also came across another paper published by Ole Miss on synthetic seeds. These same scientists were able to develop a formulation that consistently would regenerate 100% of these synthetic seeds under greenhouse conditions. Impressive statistics, but not an original or groundbreaking discovery; synthetic seeds have been around a while and are underutilized technology.
We have also been tooling around in our spare time with synthetic seeds and have some preliminary observations on their implementation.
[caption id="attachment_590" align="aligncenter" width="225"] Our first batch of synthetic seeds.[/caption]
For the rest of this article see:
/tissue-culture/
Pharmer Joe
Our first batch of synthetic seeds.[/caption]
For the rest of this article see:
/tissue-culture/
Pharmer Joe
Sign in to set favorite
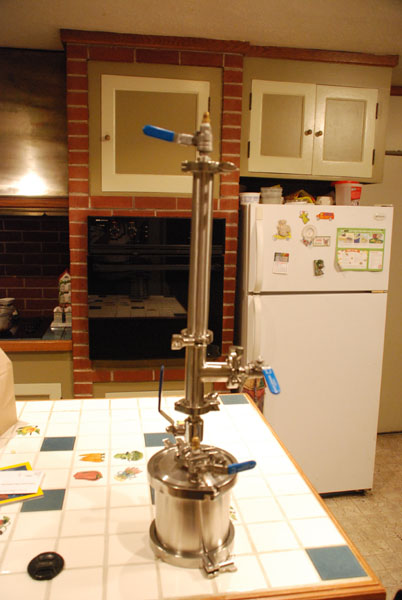





Hey there folks @ Skunk, much respect- I really enjoyed this. Being a machinist, I went to the shop and grabbed some 1" ID chrome moly tubing and some 1" OD cold rolled steel and 30 min later, had a nice pressing device for our 20 ton press. Followed your guidance, I was making nice 1/4" slugs of canna-buttons shortly after. Going to try making the various versions to test and pass around. This was a nice discovery as I have not really thought about why trim and leaf was harsh, the analogy makes perfect sense. Thanks for taking the time to present this us. Warm regards, The Captain
wonderful idea guys... love it.... now as far as trim bho goes... have yall found it works well with cooking?
Have you tried crumbling them up and adding them to food products? Seems like a half dozen in some cornbread would be delicious.
Interesting that your rooting formula only contains IBA. The study I read (on hemp) said they had better results with 0.1ppm IBA and 0.05ppm NAA. http://www.pakbs.org/pjbot/PDFs/41(2)/PJB41(2)603.pdf
marijuana from test tubes is now available on the kindle through amazon
Be sure to stop by the Skunk Pharm Research Facebook page and your likes.:) https://www.facebook.com/SkunkPharm
Great ideas, but... Not sure that I'd be keen on tasting the product from those pic's, since it looks like you're using a high LEAD content steel cap end (Lead added to prevent sparks I think, like your petrol/gas pump fittings at the petrol/gas stations). If this is the case it would be virtually impossible to avoid traces of lead containing metal shavings/dust entering your product and then probably entering your body (yes its volatile with many cases of poisoning being documented from lead fumes). Not good considering lead builds up in your body, though the traces (so far) PROBABLY wouldn't have added more lead into your system than you had gained during those bad ol' days of leaded petrol/gas resulting in lead being spewed into the air we breath and its surrounds. At least times are improving and we learn from our mistakes (if we're smart & lead dints our I.Q.!). It's worth noting here too that lead is found in high concentrations in plastic electrical insulation surrounding wires (up to 4% w/w) and in even higher concentrations in many common flexible wires and metals (it's added to make them softer), and that this lead is absorble through skin and lungs as well as things like acidic nutrient solutions, acid rain (which it always is) etc. - the less lead near you the better!
Neither the Black Iron Pipe, The carbon steel pin, nor the 304 SS used for this purpose is high in lead. Where did you get the idea that it was?
This information is invaluable. Where can I find out more?
I would suggest contacting joeoakes@skunkpharmresearch.com and picking his brain or attending one of our classes http://skunkpharmresearch.com/skunk-pharm-research-classes/
soylent green ?
Hee, hee, hee........... Cannabuttons and Super Cannabuttons are certainly green and a way to medicate more people with less. Not unlike the movie using recycle to feed the masses.
Heeeheee.... you read my mind! GMTA!
This is what I was referring to. Identification of candidate genes affecting Δ9-tetrahydrocannabinol biosynthesis in Cannabis sativa http://jxb.oxfordjournals.org/content/60/13/3715.full
This is quite fantastic and right down my alley (im a senior year biotech major). I am also part of the largest NORML chapter (UCF), how should i go about getting funding for you all? also, cannabulent could you link me to the site discussing the mRNA profiles?
Hey Ryan, We are not a registered nonprofit nor are we looking to sell any ownership stake. So any funding would have to be an in kind gift. I believe the study that cannabulent referred to can be found at the nih.gov site. There you can find the est's related to your search. Joe
Yup, qRT-PCR would work really well, in a 96 well or 384 well format, you could look at transcript levels for every gene in the pathway at once! It would be super interesting to see how distinct chemotypes (CBD vs. THC rich for example) would differ. And as deep sequencing gets cheaper, it will get easier and more quantifiable to get this type of data. With regards to mutations that affect flower morphology, as a proof of principle for the genetic transformation, it could be interesting to make GFP flowers. As the marketplace saturates, one could make quite a splash with glow in the dark flowers. Also doing traditional forward genetics with chemical mutagens could be a lot of fun.
I will leave the genome sequence to Medicinal Genomics since they already have an Ion Torrent sequencer and I don't have an extra 50k. Gfp is a fun marker to use and its not the only glowing gene. Check out Invitrogens living rainbow collection. You could use them in concert with different promoters for different localized expression, like green trichomes and red flowers. Joe
Yes, I am familiar with the latter study, but it was my understanding that once transformed, the researchers were unable to fully differentiate the plant into a vegetative state. To that point, do you have any pictures of older clones, ie. ones that look more well differentiated? In regards to the mRNA expression profiling. This has been done for cannabis trichomes, so the protocol and sequences are out there, using academic resources one could do this pretty easily. I like your idea of looking at this as a function of development, it would also be interesting to look across sub-types. These are experiments that would be foundational to any attempts to engineer and/or disrupt the cannabinoid biosynthetic pathway. One might also want to look at the GC-MS at the same time and then try to correlate the transcriptional changes with the chemical changes. until next time!
I know that they have done some Mrna studies and that they have found the specific genes I'm interested in. I was thinking120 clones and analyzing one at each change in the photoperiod. 120 because I would destroy each specimen to do the analysis. I would really like an rtpcr for Christmas but alas I'm not that good of a boy. That would be more interesting to me than the current studies. I'm also interested in mutations that affect flower morphology. I look forward to your comments. Joe
I am really impressed with the site and was very excited by this page. Are you mostly pursuing this type of work so that you can store genetic copies/clones of a given strain? It is my understanding that there are no publicly documented cases for the genetic transformation of cannabis. I fantasize about doing so. One could use trichome specific promoters to manipulate terpene and cannabinoid diversity and concentration. But what I would really like to do, is to play with the genes in the trichome development pathway. Work in Arabidopsis has led to the discovery of genes that dictate size, concentration and cellularity of trichome structure. Knocking out homologous repressors of trichome growth in cannabis could be very interesting and could serve as a model system for the use of a plant as a factory for the production of novel/difficult to synthesize natural products. Keep up the good work! You guys rock! (PS: I do not work for Monsanto, nor would I endorse the industrialization of said engineered strains)
Hello Cannabulent, In response to your knowlegeable comment: We are using TC to store clones for future genetic comparison. Our DNA project is currently on back burner status because it’s expensive. We aren't currently doing any in vitro DNA assays; that doesn’t mean we aren't paying attention to literature published that helps our on paper project outline. In the future we hope to be able link mutations in the cannabinoid biosynthetic pathway to GC analysis of the trichomes. I am particularly interested in gene expression analysis of the floral bracts and trichomes as they develop. Soon we will be adding a page about our DNA aspirations so keep an eye out for it. http://www.medicinalgenomics.com (Whole Genome Sequencing) There is one published report of transformation that I am aware of (see link) I think they would have been more successful obtaining transgenic plants with a floral dip while the seeds are still immature. Transformation http://www.jstor.org/discover/10.2307/4293671?uid=3739856&uid=2129&uid=2&uid=70&uid=4&uid=3739256&sid=47698763684267 Joe